Code (GitHub) | Paper (bioRxiv)
MatBots are primitive AIs, 'assistants' if you will, that use minimalistic GUI dialogs to guide the user through a data processing pipeline in Matlab.
Isn't that an 'app'? Bots are much more restrictive than apps. Users are, to a greater extent than in an app, guided through the correct steps to perform a task. A bot usually performs a much more limited task than an app.
When possible, bots have a 'headless' mode, which allows them to execute a processing pipeline as a typical Matlab function, either on an image or a folder of images.
The first crop of bots perform selected biological-image-analysis pipelines, such as nuclei segmentation and point-source scoring. Source code and more details are available on GitHub. Below is a list of currently implemented bots, with links to video tutorials and sample datasets when available.
For even more details, and to cite this work, please refer to the bioRxiv manuscript.
Clone the GitHub repository to access all bots and tools. Individual, self-contained bots and tools can be downloaded from File Exchange.
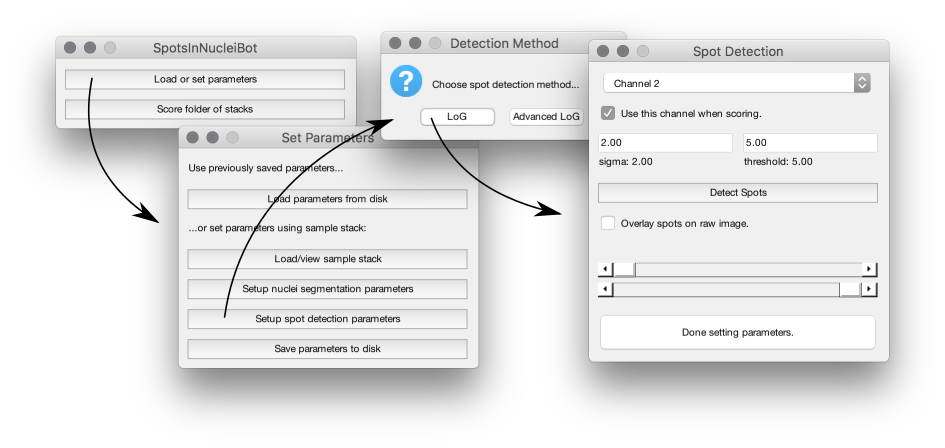
A bot to count point sources (spots) in nuclei.
Assists in training a Machine Learning model to segment nuclei.
For Machine Learning experts: the source code to train a stacked Random Forest for nuclei segmentation, using circularity features, is available here. This algorithm is used as the back-end for pixel classification in NucleiSegmentationBot.
Assists in training a Machine Learning model to segment grayscale images. This is a more general implementation of NucleiSegmentationBot.
A stand-alone bot to annotate images (the same used as a module in PixelClassificationBot and NucleiSegmentationBot).
Analogous to ImageAnnotationBot for 3D volumes.
A bot to make stacks out of image planes. This can be used to assemble stacks for SpotsInNucleiBot.
A bot to measure the size of spot-like (including diffraction-limited) objects.
Assists in segmenting objects using a simple 3-step pipeline consisting of (1) smoothing, (2) thresholding, and (3) watershed on distance transform. Allows post-processing operations to filter masks based on area/eccentricity, and grow or shrink objects by a constant amount.
For questions and feedback, contact Marcelo.