The Image and Data Analysis Core and the Office for Academic and Research Integrity at Harvard Medical School have an ongoing collaboration developing tools to identify re-use of images in academic publications. To tackle this problem at scale, our team is currently developing methods to determine if, given two images, one is a manipulated version of the other -- and, in case they are, further identify subregions in the images that provide matching evidence. Manipulations include a broad range of transformations representing common alterations and obfuscations ranging from simple crop and rigid transformation to projective and nonlinear transformations, histogram adjustment, or local alterations such as partial erasing, the addition of text and annotations, etc. We therefore apply metric-learning approaches based on convolutional neural networks to produce similarity functions which are invariant to these manipulations. These models are trained on a large database of synthetically altered biological images and tested on real-world cases of image manipulation which were discovered in peer-reviewed publications.
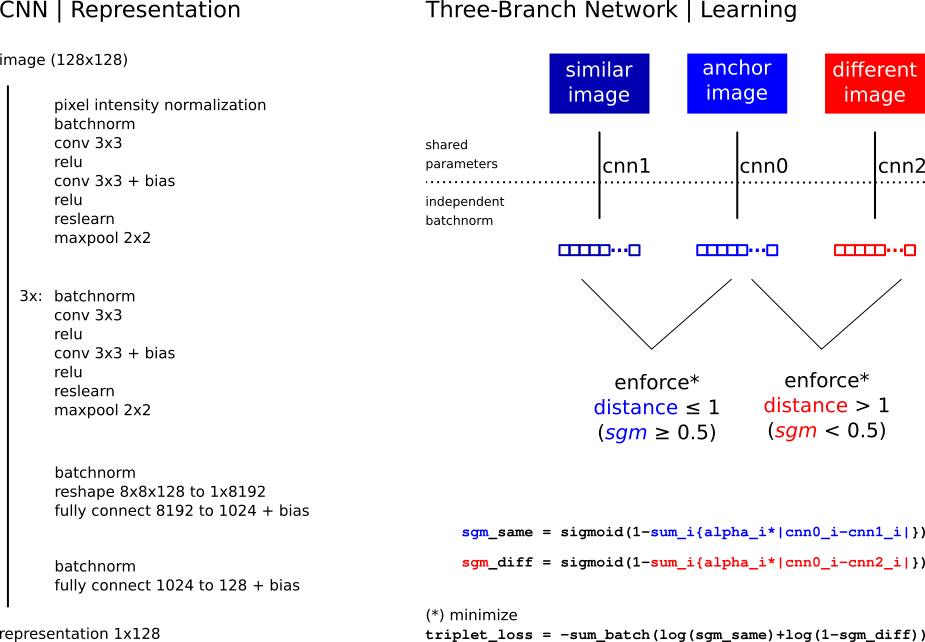
In the first phase of this project, we implemented a siamese network model for duplicate detection, and curated a dataset of synthetic images for training.
Code (GitHub) | Paper (arXiv) | Database (Dataverse)
In the second phase, we further tested a range of deep learning models for duplicate detection, and curated a substantially larger dataset of synthetic images. We are currently working on methods to align (register) images once they're flagged as duplicates, and on methods to quantify similarity of overlapping areas (after registration).
BINDER: Bio-Image Near-Duplicate Examples Repository
Last updated Jun 23 2021.